So when we analyze such patterns to make inferences about the underlying processes, the results of the analysis needs to be interpreted differently for polyploids than for diploids. In polyploids, these processes work differently, resulting in different patterns in the distribution of genetic variation. Therefore, the action of evolution depends heavily on the processes that influence the distribution of genetic variation within species, most importantly genetic drift, breeding system, mode of inheritance, segregation patterns, migration, mutation, and selection.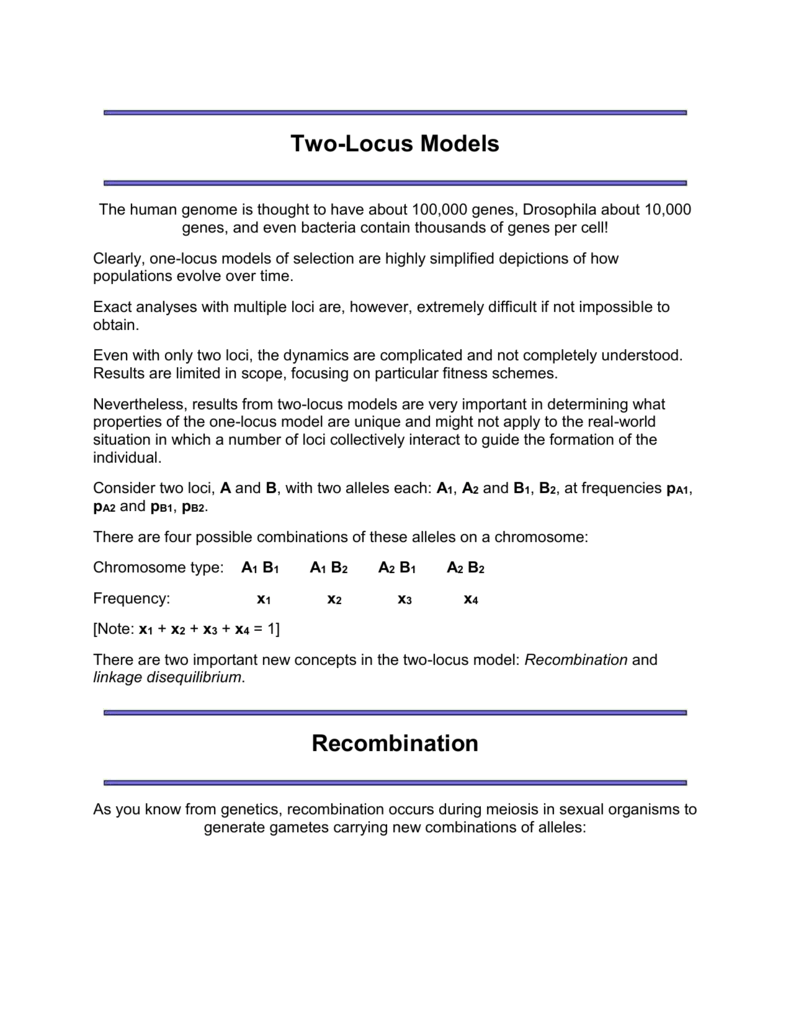  
2014 Parisod and Broennimann 2016).Īt its most basic, evolution is simply a shift in allele frequencies. 2008) and ecological preferences ( Glennon et al. 2017) and may differ in life-history characteristics, such as mating system ( Meirmans et al. These cytotypes may show striking geographic distributions ( Suda et al. Furthermore, there are many species-especially in plants-in which multiple cytotypes are present. Polyploidy also plays a direct role in the speciation process: an estimated 2–4% of speciation events in flowering plants and 7% in ferns are thought to be the result of polyploidization ( Otto and Whitton 2000). Even the genome of Arabidopsis thaliana, one of the smallest genomes among the angiosperms, shows evidence for several rounds of whole genome duplication ( Bomblies and Madlung 2014). Phylogeography, population structure, autopolyploidy, genetic differentiation, genetic diversity, Hardy–Weinberg, population structure, tetrasomyĭuring their evolution, many plant and animal taxa have gone through one or several rounds of genome duplication ( Ramsey and Schemske 1998). Therefore, simulations such as we used throughout this review are an important tool to verify the results of analyses of polyploid genetic data. Furthermore, the availability of more data may aggravate the biases that can arise, and increase the risk of false inferences. Modern sequencing techniques will soon be able to overcome some of the current limitations to the analysis of polyploid data, though the techniques are lagging behind those available for diploids. From our overview, it is clear that the statistical toolbox that is available for the analysis of genetic data is flexible and still expanding. We also discuss for each type of inference what biases may arise from the polyploid-specific complications and how these biases can be overcome. For each, we point out how the statistical approach, expected result, and interpretation differ between different ploidy levels. We discuss several widely used types of inferences, including genetic diversity, Hardy–Weinberg equilibrium, population differentiation, genetic distance, and detecting population structure. Here, we review the theoretical and statistical aspects of the population genetics of polyploids. This is because of several polyploidy-specific complications in segregation and genotyping such as tetrasomy, double reduction, and missing dosage information. This is unfortunate since the analysis of polyploid genetic data-and the interpretation of the results-requires even more scrutiny than with diploid data. Though polyploidy is an important aspect of the evolutionary genetics of both plants and animals, the development of population genetic theory of polyploids has seriously lagged behind that of diploids.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
